Evaluation of Machine Learning and Deep Learning Approaches for Automatic Detection of Eye Diseases
DOI:
https://doi.org/10.58190/imiens.2024.85Keywords:
Classification, Deep features, Deep learning, Eye diseases, Machine learningAbstract
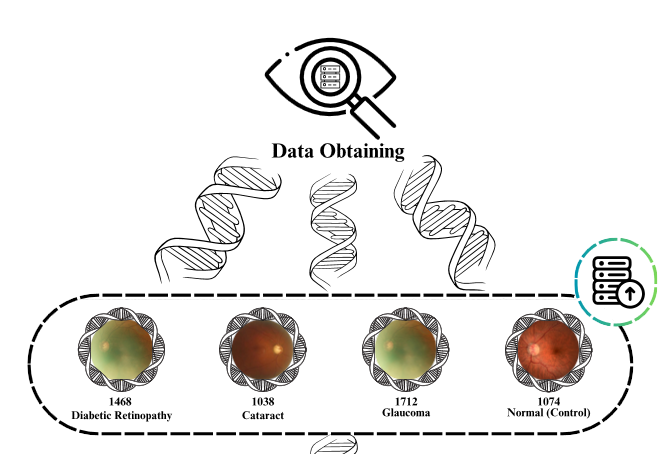
There are many ocular diseases present in the world. These diseases may arise from factors such as genetic predisposition, environmental influences, and aging. In recent years, advancements in technology have facilitated the detection of ocular pathologies through machine learning techniques. Machine learning models can serve as decision support mechanisms in diagnostic scenarios. In this study, the aim is to detect ocular diseases using machine learning and deep learning techniques. To enhance the results obtained from classification with 4,217 images in the study, 705 images were added to the glaucoma class and 370 images were added to the Diabetic Retinopathy class. The supplemented dataset with additional images comprises a total of four classes. One class represents the control group and is labeled as "normal." The remaining three classes represent disease categories: Diabetic Retinopathy, Cataract, and Glaucoma. To extract deep features from the images, a pre-trained InceptionV3 model was utilized, resulting in 2048 features extracted. These extracted features were then classified using Neural Network (NN), Logistic Regression (LR), k-Nearest Neighbors (k-NN), and Random Forest (RF) machine learning models. Before the dataset supplemented with additional images, the classification accuracies of the machine learning models were as NN 89.2%, LR 87.3%, k-NN 81.2%, and Random Forest 76.9%. Upon examining the classification accuracies after dataset supplemented with additional images, the following improvements were observed: NN 90.9% with a 1.7% increase, LR 90.2% with a 2.9% increase, k-NN 84.6% with a 3.4% increase, and Random Forest 82% with a 5.1% increase. Performance evaluation was conducted using recall, precision, and F-1 score metrics. Additionally, the learning performance of the machine learning models was assessed through Receiver Operating Characteristic (ROC) curves and Area Under Curve (AUC) values.
Downloads
References
Raza, A., et al., Classification of Eye Diseases and Detection of Cataract using Digital Fundus Imaging (DFI) and Inception-V4 Deep Learning Model, in 2021 International Conference on Frontiers of Information Technology (FIT). 2021. p. 137-142. DOI: 10.1109/FIT53504.2021.00034
Umesh, L., M. Mrunalini, and S. Shinde, Review of image processing and machine learning techniques for eye disease detection and classification. International Research Journal of Engineering and Technology, 2016. 3(3): p. 547-551.
Pan, Y., K. Zhao, and Z. Tan, Fundus image classification using Inception V3 and ResNet-50 for the early diagnostics of fundus diseases. Frontiers in Physiology, 2023. 14: p. 1126780. DOI: 10.3389/fphys.2023.1126780
Bitto, A.K. and I. Mahmud, Multi categorical of common eye disease detect using convolutional neural network: a transfer learning approach. Bulletin of Electrical Engineering and Informatics, 2022. 11(4): p. 2378-2387. DOI: 10.11591/eei.v11i4.3834
Hameed, S.R. and H.M. Ahmed, Eye diseases classification using back propagation with parabola learning rate. Al-Qadisiyah Journal Of Pure Science, 2021. 26(1): p. 1–9-1–9. DOI: 10.29350/qjps2021.26.1.1220
Smaida, M., S. Yaroshchak, and Y. El Barg. DCGAN for Enhancing Eye Diseases Classification. in CMIS. 2021.
Ahmad, H.M. and S.R. Hameed. Eye Diseases Classification Using Hierarchical MultiLabel Artificial Neural Network. IEEE. DOI: 10.1109/IT-ELA50150.2020.9253120
Wang, W., et al., Learning Two-Stream CNN for Multi-Modal Age-Related Macular Degeneration Categorization. IEEE Journal of Biomedical and Health Informatics, 2022. 26(8): p. 4111-4122. DOI:10.1109/JBHI.2022.3171523
Hirota, M., et al., Effect of color information on the diagnostic performance of glaucoma in deep learning using few fundus images. International Ophthalmology, 2020. 40(11): p. 3013-3022. DOI:10.1007/s10792-020-01485-3
Sayres, R., et al., Using a deep learning algorithm and integrated gradients explanation to assist grading for diabetic retinopathy. Ophthalmology, 2019. 126(4): p. 552-564. DOI: 10.1016/j.ophtha.2018.11.016
Serener, A. and S. Serte. Transfer learning for early and advanced glaucoma detection with convolutional neural networks. in 2019 Medical technologies congress (TIPTEKNO). 2019. IEEE. DOI: 10.1109/TIPTEKNO.2019.8894965
Islam, M.T., et al. Source and camera independent ophthalmic disease recognition from fundus image using neural network. in 2019 IEEE International Conference on Signal Processing, Information, Communication & Systems (SPICSCON). 2019. IEEE. DOI: 10.1109/SPICSCON48833.2019.9065162
Lam, C., et al., Automated detection of diabetic retinopathy using deep learning. AMIA summits on translational science proceedings, 2018. 2018: p. 147.
Wang, X., et al. Diabetic retinopathy stage classification using convolutional neural networks. in 2018 IEEE International Conference on Information Reuse and Integration (IRI). 2018. IEEE. DOI: 10.1109/IRI.2018.00074
Chen, H., et al. Detection of Diabetic Retinopathy using Deep Neural Network. in 2018 IEEE 23rd International Conference on Digital Signal Processing (DSP). 2018. DOI: 10.1109/ICDSP.2018.8631882
Eye Diseases Classification Dataset. Available from: https://www.kaggle.com/datasets/gunavenkatdoddi/eye-diseases-classification.
Glaucoma Detection Dataset. Available from: https://www.kaggle.com/datasets/sshikamaru/glaucoma-detection.
Diabetic Retinopathy 224x224 Gaussian Filtered Dataset, Available from: https://www.kaggle.com/datasets/sovitrath/diabetic-retinopathy-224x224-gaussian-filtered.
Cruyff, M.J., et al., A review of regression procedures for randomized response data, including univariate and multivariate logistic regression, the proportional odds model and item response model, and self-protective responses. Handbook of statistics, 2016. 34: p. 287-315. DOI: 10.1016/bs.host.2016.01.016
Akhgarjand, C., et al., Does Ashwagandha supplementation have a beneficial effect on the management of anxiety and stress? A systematic review and meta‐analysis of randomized controlled trials. Phytotherapy Research, 2022. 36(11): p. 4115-4124. DOI: 10.1002/ptr.7598
Cinar, I. and M. Koklu, Classification of rice varieties using artificial intelligence methods. International Journal of Intelligent Systems and Applications in Engineering, 2019. 7(3): p. 188-194. DOI: 10.18201/ijisae.2019355381
Wang, D., et al. Neural networks are more productive teachers than human raters: Active mixup for data-efficient knowledge distillation from a blackbox model. in Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition. 2020.
Brownlee, J., What is the Difference Between a Batch and an Epoch in a Neural Network. Machine learning mastery, 2018. 20.
Chicco, D., Siamese neural networks: An overview. Artificial neural networks, 2021: p. 73-94. DOI: 10.1007/978-1-0716-0826-5_3
Cinar, I. and M. Koklu, Identification of Rice Varieties Using Machine Learning Algorithms. Journal of Agricultural Sciences, 2022: p. 9-9. DOI: 10.15832/ankutbd.862482
Koklu, M. and K. Sabanci, Estimation of credit card customers payment status by using kNN and MLP. International Journal of Intelligent Systems and Applications in Engineering, 2016. 4(Special Issue-1): p. 249-251.
Mao, W. and F. Wang, New advances in intelligence and security informatics. 2012: Academic Press.
Dunham, K., Chapter 6-Phishing, SMishing, and Vishing. Mobile Malware Attacks and Defense, 2009: p. 125-196.
Isik, M., Ozulku, B., Kursun, R., Taspinar, Y. S., Cinar, I., Yasin, E. T., & Koklu, M. (2024). Automated classification of hand-woven and machine-woven carpets based on morphological features using machine learning algorithms. The Journal of The Textile Institute, p. 1-10. DOI: 10.1080/00405000.2024.2309694
Singh, D., et al., Classification and analysis of pistachio species with pre-trained deep learning models. Electronics, 2022. 11(7): p. 981. DOI: 10.3390/electronics11070981
Koklu, M., et al., A CNN-SVM study based on selected deep features for grapevine leaves classification. Measurement, 2022. 188: p. 110425. DOI: 10.1016/j.measurement.2021.110425
Koklu, M., I. Cinar, and Y.S. Taspinar, CNN-based bi-directional and directional long-short term memory network for determination of face mask. Biomedical signal processing and control, 2022. 71: p. 103216. DOI: 10.1016/j.bspc.2021.103216
Demir, A., F. Yilmaz, and O. Kose. Early detection of skin cancer using deep learning architectures: resnet-101 and inception-v3. in 2019 medical technologies congress (TIPTEKNO). 2019. IEEE. DOI: 10.1109/TIPTEKNO47231.2019.8972045
Sinha, R. and J. Clarke. When technology meets technology: Retrained ‘Inception V3’classifier for NGS based pathogen detection. in 2017 IEEE International Conference on Bioinformatics and Biomedicine (BIBM). 2017. IEEE. DOI: 10.1109/BIBM.2017.8217942
Szegedy, C., Vanhoucke, V., Ioffe, S., Shlens, J., & Wojna, Z. (2016). Rethinking the inception architecture for computer vision. In Proceedings of the IEEE conference on computer vision and pattern recognition (pp. 2818-2826).
Çataloluk, H., Gerçek tıbbi veriler üzerinde veri madenciliği yöntemlerini kullanarak hastalık teşhisi. 2012, Bilecik Üniversitesi, Fen Bilimleri Enstitüsü.
Ruuska, S., et al., Evaluation of the confusion matrix method in the validation of an automated system for measuring feeding behaviour of cattle. Behavioural processes, 2018. 148: p. 56-62. DOI: 10.1016/j.beproc.2018.01.004
Gencturk, B., Arsoy, S., Taspinar, Y. S., Cinar, I., Kursun, R., Yasin, E. T., and Koklu, M. (2024). Detection of hazelnut varieties and development of mobile application with CNN data fusion feature reduction-based models. European Food Research and Technology, 250(1), p. 97-110. DOI: 10.1007/s00217-023-04369-9
Koklu, M. and I.A. Ozkan, Multiclass classification of dry beans using computer vision and machine learning techniques. Computers and Electronics in Agriculture, 2020. 174: p. 105507.
Browne, M.W., Cross-validation methods. Journal of mathematical psychology, 2000. 44(1): p. 108-132. DOI: 10.1016/j.compag.2020.105507
Bates, S., T. Hastie, and R. Tibshirani, Cross-validation: what does it estimate and how well does it do it? Journal of the American Statistical Association, 2023: p. 1-12. DOI: 10.1080/01621459.2023.2197686
Xiong, Z., et al., Evaluating explorative prediction power of machine learning algorithms for materials discovery using k-fold forward cross-validation. Computational Materials Science, 2020. 171: p. 109203. DOI: 10.1016/j.commatsci.2019.109203
Maleki, F., et al., Machine learning algorithm validation: from essentials to advanced applications and implications for regulatory certification and deployment. Neuroimaging Clinics, 2020. 30(4): p. 433-445. DOI: 10.1016/j.nic.2020.08.004
Downloads
Published
Issue
Section
License
Copyright (c) 2024 Intelligent Methods In Engineering Sciences

This work is licensed under a Creative Commons Attribution-ShareAlike 4.0 International License.






